[ad_1]
Study population
The algorithms were evaluated on the publicly available MIMIC-III data set (ethics approval was not required)32. This data set comprises deidentified health data associated with 61,532 ICU stays and 46,476 critical care patients at Beth Israel Deaconess Medical Center in Boston, Massachusetts between 2001 and 2012.
The supervised learning task consists of predicting, for a given ICU stay, whether the patient will be readmitted to the ICU within 30 days from discharge. Patients were excluded if they died during the ICU stay (N = 4,787 ICU stays), were not adults (18 years old or older) at the time of discharge (N = 8,129 ICU stays) or died within 30 days from discharge without being readmitted to the ICU (N = 3,318 ICU stays). The final data set comprised 45,298 ICU stays for 33,150 patients, labelled as either positive (N = 5,495) or negative (N = 39,803) depending on whether a patient did or did not experience readmission within 30 days from discharge. To develop and evaluate the algorithms, patients were subdivided randomly into training and validation (90%) and test sets (10%). This subdivision was based on patient identifiers and not on ICU stay identifiers to prevent information leaks between data sets (since the prediction is based on the entire clinical history of a patient).
Model variables
The EMR of a patient can be represented as a set of static variables and timestamped codes. In the present study, static variables included the patient’s gender, age, ethnicity, insurance type, marital status, the previous location of the patient prior to arriving at the hospital (admission location), and whether the patient was admitted for elective surgery. Both length of ICU stay and length of hospital stay prior to ICU admission were recorded. An additional static variable was given by the number of ICU admissions in the year preceding the considered index ICU stay.
Data types of timestamped codes included international classification of diseases and related health problems (ICD-9) diagnosis and procedure codes, prescribed medications, and patient vital signs. All diagnosis and procedure codes in the clinical history of a patient were considered for predictive purposes; however, prescribed medications and recorded vital signs were restricted to the ICU stay of interest. Following the OASIS severity of illness score29, assessed vital signs comprised the Glasgow Coma Scale score (sum of eye response, verbal response, motor response components), heart rate, mean arterial pressure, respiratory rate, body temperature, urine output, and whether the patient necessitated ventilation. Continuous measurements of vital signs were categorised in the same manner as in OASIS and assigned corresponding codes29. To reduce redundant information, whenever the same vital sign-related code was recorded consecutively more than once, only the latest observation was kept in the data.
Elapsed times, measured in days, associated with diagnosis and procedure codes were based on the date and time of discharge from the corresponding hospital admission. Elapsed times, measured in hours, associated with medications and vital signs were based on the date and time of prescription start and measurement, respectively. In the present study, the simplifying assumption is made that diagnosis and procedure codes are available immediately at the time of discharge from the ICU. Categorical values of static variables or timestamped codes associated with less than 100 ICU stays were re-labelled as “other”.
Artificial neural network architectures
Several “deep” neural network architectures for predicting a patient’s risk of readmission to the ICU were implemented and compared. To make this comparison as fair as possible, all architectures shared a similar high level structure: (1) timestamped codes were mapped to vector embeddings; (2) numerical scores associated with diagnosis and procedure codes, and with medication and vital sign codes, were computed using attention mechanisms and/or recurrent layers; (3) these scores were concatenated with the static variables and passed on to a “logistic regression layer” (i.e. a fully connected layer with a sigmoid activation function). Further details about individual network components are reported in the following sections.
Embeddings
Diagnosis and procedure codes, as well as medication and vital sign codes, were mapped to corresponding “embeddings” (real-valued vectors). The size of these embeddings was set proportional to the fourth root of the total number of codes in the dictionary (diagnoses/procedures and medications/vital signs were processed separately since they were measured on different time scales)7. Time-aware code embeddings were computed in three different manners. A first approach used MCEs with time-aware attention19. MCEs are based on the continuous bag-of-words model33, but instead of using fixed-sized temporal windows to determine a code’s context, attention mechanisms learn the temporal scope of a code together with its embedding. A second approach optimised an embedding matrix at the same time as the other parameters of the network and, optionally, concatenated the elapsed times to the resulting vectors. A third approach optimised an embedding matrix at the same time as the other parameters of the network and modelled the dynamics in time of the computed embeddings using neural ODEs16,17,18. More in detail, the embedding of a code at time zero (i.e. at the time of discharge from the ICU) was stored in the embedding matrix whereas the embedding of a code recorded before discharge was computed by solving an initial value problem where derivatives with respect to time were approximated by a multilayer perceptron.
Attention and/or recurrent layers
The sequence of code embeddings associated with a patient is usually of arbitrary length and needs to be integrated into a fixed-size vector for further processing. Attention mechanisms, such as dot-product attention34, compute a weighted average of the code embeddings, where a higher weight is assigned to the most relevant codes. Alternatively, recurrent layers iteratively process an input sequence of codes and, at each iteration, update an internal memory state and generate an output vector8,9. Information may be integrated for further processing by using either the final memory state of the recurrent cell or by applying an attention mechanism to the set of output vectors5,6,7. Specifically, in this work, recurrent cells were implemented using bi-directional gated recurrent units9. Time-related information was taken into account by concatenating the time differences between observations to the embedding vectors, by applying an exponential decay proportional to the time differences between observations to the internal memory state of the recurrent cell13,14,15 or by modelling the dynamics in time of the internal memory state using neural ODEs16,17,18. To aid subsequent interpretation without altering network capacity, the fixed-size vectors produced by attention mechanisms and/or recurrent layers were reduced to two scalar-valued scores (one related to diagnoses/procedures and one related to medications/vital signs) using fully connected layers with a linear activation function.
Logistic regression layer
The computed diagnoses/procedures and medications/vital signs scores were concatenated to the vector of static variables and passed to a fully connected layer with a sigmoid activation function. The output of the network corresponds to the risk of readmission to the ICU within 30 days from discharge.
Architectures
The following neural network architectures were compared for predicting readmission to the ICU:
-
ODE + RNN + Attention: dynamics in time of embeddings are modelled using neural ODEs, embeddings are passed to RNN layers, dot-product attention is applied to RNN outputs.
-
ODE + RNN: dynamics in time of embeddings are modelled using neural ODEs, embeddings are passed to RNN layers, the final memory states are used for further processing.
-
RNN (ODE time decay) + Attention: embeddings are passed to RNN layers with dynamics in time of the internal memory states modelled using neural ODEs, dot-product attention is applied to RNN outputs.
-
RNN (ODE time decay): embeddings are passed to RNN layers with dynamics in time of the internal memory states modelled using neural ODEs, the final memory states are used for further processing.
-
RNN (exp time decay) + Attention: embeddings are passed to RNN layers with internal memory states decaying exponentially over time, dot-product attention is applied to RNN outputs.
-
RNN (exp time decay): embeddings are passed to RNN layers with internal memory states decaying exponentially over time, the final memory states are used for further processing.
-
RNN (concatenated Δtime) + Attention: embeddings are concatenated with time differences between observations and passed to RNN layers, dot-product attention is applied to RNN outputs.
-
RNN (concatenated Δtime): embeddings are concatenated with time differences between observations and passed to RNN layers, the final memory states are used for further processing.
-
ODE + Attention: dynamics in time of embeddings are modelled using neural ODEs, dot-product attention is applied to the embeddings.
-
Attention (concatenated time): embeddings are concatenated with elapsed times, dot-product attention is applied to the embeddings.
-
MCE + RNN + Attention: MCE is used to compute the embeddings, embeddings are passed to RNN layers, dot-product attention is applied to RNN outputs.
-
MCE + RNN: MCE is used to compute the embeddings, embeddings are passed to RNN layers, the final memory states are used for further processing.
-
MCE + Attention: MCE is used to compute the embeddings, dot-product attention is applied to the embeddings.
The dimension of the internal memory state of RNN cells was set equal to the dimension of the input embeddings. Similarly, the dimension of the hidden representation of embeddings when computing dot-product attention was left unchanged. Derivatives with respect to time used to implement neural ODEs were approximated by a multilayer perceptron with three hidden layers of constant width equal to the size of the input. The Euler method was used as ODE solver.
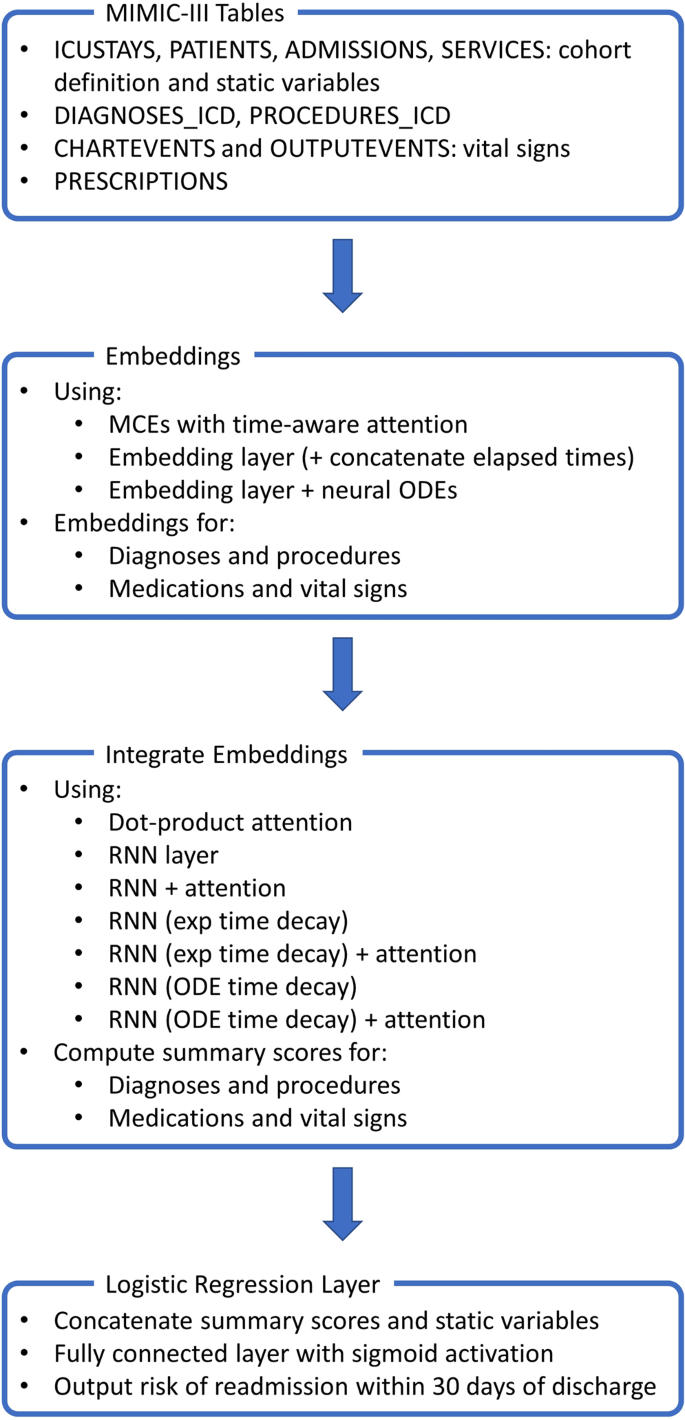
An overview of the considered neural network architectures is presented in Fig. 1. For completeness, the deep learning approaches were also compared with a logistic regression model using all static variables and the most recent vital signs for each patient as covariates.
Interpretation of attention-based models
For the proposed neural network architectures, the weights of the final fully connected layer can be used to determine the impact of static variables and timestamped codes on estimated risk. As in traditional logistic regression, these weights can be interpreted as increases in log-odds for unplanned early ICU readmission if the corresponding static variables or scores are increased by one unit.
It is also of interest to determine which codes (i.e. diagnoses, procedures, medications, vital signs) are associated with a prediction of high risk. Dot-product attention computes a weighted average of embedded codes; fully connected layers are then used to output scores associated with diagnoses/procedures and medications/vital signs. By passing single codes (i.e. the rows of the embedding matrix) to the fully connected layers computing these scores, it is possible to associate each code with a score. The higher the score, the higher the risk of ICU readmission when a patient’s EMR contains that code.
To estimate Bayesian credible intervals around network weights and computed risk scores, the posterior distribution of weights was approximated using stochastic variational inference with mean-field approximation35,36. In the present study, the variational posterior is assumed to be a diagonal Gaussian distribution and is estimated using the Bayes by Backprop algorithm37. Following the original paper, a priori sparsity of the network weights is encouraged by formulating the prior distribution as a scale mixture of two zero-mean Gaussian densities with standard deviations of σ1 = 1 and σ2 = e−6, respectively, and mixture weight π = 0.537. After the posterior distribution has been computed, 95% credible intervals around network weights (or combinations thereof) can be estimated by repeated sampling. Sampling of network weights may also be used to compute credible intervals around the risk prediction for a given patient.
Training
To compare the classification accuracy of the considered neural network architectures, maximum likelihood estimates of network parameters were obtained using a log-loss cost function on the training data, extensive use of dropout with 50% probability after each embedding, RNN, and attention layers38, and stochastic gradient descent with an Adam optimizer (batch size of 128 and learning rate of 0.001)39. Class imbalance was taken into consideration by assigning a proportionally higher cost of misclassification to the minority class40. Training was terminated after 80 epochs since overfitting of the training data started to become apparent with additional training epochs (based on average precision on the validation data). For interpretation purposes, Bayes by Backprop was used to train the “Attention (concatenated time)” neural network architecture on the entire data set, terminating if the loss function (the expected lower bound) did not decrease for 10 consecutive epochs.
Statistical analysis
Baseline characteristics were determined for the analysed patient population. The prediction accuracy of each considered algorithm was evaluated based on average precision, AUROC, F1-Score, sensitivity, and specificity. Average precision may reflect algorithmic performance on imbalanced data sets better than AUROC as it does not reward true negatives41,42. The F1-Score was maximised over different threshold values on risk predictions. Sensitivity and specificity were computed by maximising Youden’s J statistic43. 95% confidence intervals associated with each metric were computed by bootstrapping, i.e. by sampling the test set with replacement 100 times and re-evaluating the models each time44. Since the bootstrap estimator assumes the resampling of independent events, sampling was based on patient identifiers rather than on ICU stay identifiers.
Training the “Attention (concatenated time)” network using Bayes by Backprop allowed computation of odds ratios (OR) associated with static variables and ranking of the timestamped codes (diagnoses, procedures, medications, and vital signs) according to their associated average scores (a high positive score corresponds to increased risk of readmission to the ICU); corresponding 95% credible intervals were determined using 10,000 network samples.
Software was implemented in Python using Scikit-learn45 and PyTorch46; the developed algorithms are publicly available at https://github.com/sebbarb/time_aware_attention.



![Buildbox Free - How To Make 2D Platformer Game [PART 1]](https://e928cfdc7rs.exactdn.com/info/uploads/sites/3/2020/01/Buildbox-Free-How-To-Make-2D-Platformer-Game-PART-150x150.jpg?strip=all)

